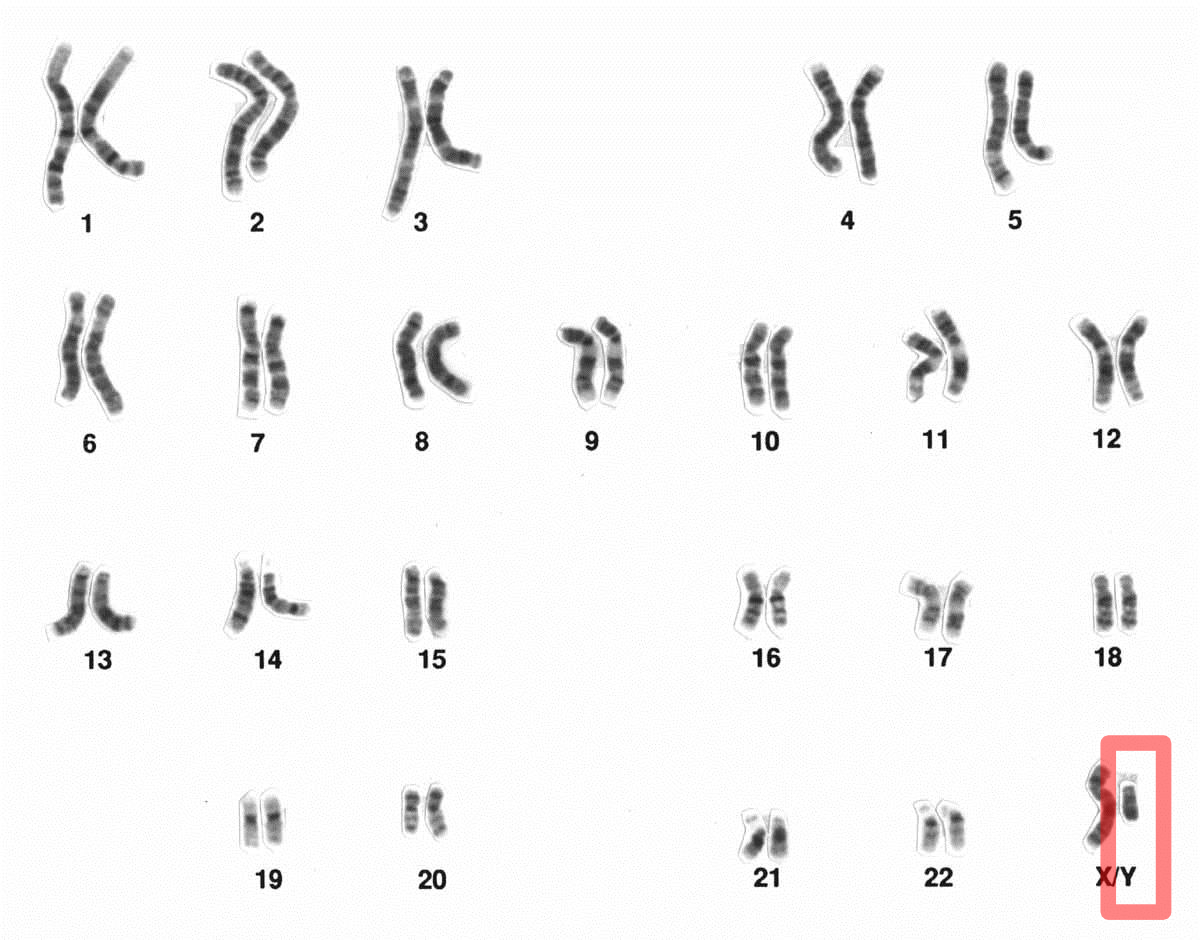
A síndrome de coroideremia-surdez-obesidade é uma distrofia retiniana ligada ao X caracterizada por coroideremia, causando nos homens afetados nictalopia progressiva e eventual cegueira central. Obesidade, deficiência intelectual moderada e surdez congênita mista (sensorial e condutiva) também são observadas. Portadoras do sexo feminino apresentam alterações retinianas típicas indicativas do estado de portador de coroideremia.
Introdução
O que você precisa saber de cara
A síndrome de coroideremia-surdez-obesidade é uma distrofia retiniana ligada ao X caracterizada por coroideremia, causando nos homens afetados nictalopia progressiva e eventual cegueira central. Obesidade, deficiência intelectual moderada e surdez congênita mista (sensorial e condutiva) também são observadas. Portadoras do sexo feminino apresentam alterações retinianas típicas indicativas do estado de portador de coroideremia.
Escala de raridade
<1/50kMuito rara
1/20kRara
1/10kPouco freq.
1/5kIncomum
1/2k
Encontrou um erro ou informação desatualizada? Sugira uma correção →
Entender a doença
Do básico ao detalhe, leia no seu ritmo
Preparando trilha educativa...
Sinais e sintomas
O que aparece no corpo e com que frequência cada sintoma acontece
Partes do corpo afetadas
+ 12 sintomas em outras categorias
Características mais comuns
Os sintomas variam de pessoa para pessoa. Abaixo estão as 49 características clínicas mais associadas, ordenadas por frequência.
Linha do tempo da pesquisa
Encontrou um erro ou informação desatualizada? Sugira uma correção →
Genética e causas
O que está alterado no DNA e como passa nas famílias
Genes associados
1 gene identificado com associação a esta condição. Padrão de herança: X-linked recessive.
Probable transcription factor which exert its primary action widely during early neural development and in a very limited set of neurons in the mature brain
Nucleus
Deafness, X-linked, 2
A form of deafness characterized by both conductive hearing loss resulting from stapes (perilymphatic gusher) fixation, and progressive sensorineural deafness.
Variantes genéticas (ClinVar)
274 variantes patogênicas registradas no ClinVar.
Diagnóstico
Os sinais que médicos procuram e os exames que confirmam
Tratamento e manejo
Remédios, cuidados de apoio e o que precisa acompanhar
Onde tratar no SUS
Hospitais de referência no Brasil e o protocolo oficial do SUS (PCDT)
🇧🇷 Atendimento SUS — Síndrome de microdeleção Xq21
Selecione um estado ou use sua localização para ver resultados.
Dados de DATASUS/CNES, SBGM, ABNeuro e Ministério da Saúde. Sempre confirme a disponibilidade diretamente com o estabelecimento.
Pesquisa ativa
Ensaios clínicos abertos e novidades científicas recentes
Pesquisa e ensaios clínicos
Nenhum ensaio clínico registrado para esta condição.
Publicações mais relevantes
Contiguous Gene Syndromes and Hearing Loss: A Clinical Report of Xq21 Deletion and Comprehensive Literature Review.
Given the crucial role of the personalized management and treatment of hearing loss (HL), etiological investigations are performed early on, and genetic analysis significantly contributes to the determination of most syndromic and nonsyndromic HL cases. Knowing hundreds of syndromic associations with HL, little comprehensive data about HL in genomic disorders due to microdeletion or microduplications of contiguous genes is available. Together with the description of a new patient with a novel 3.7 Mb deletion of the Xq21 critical locus, we propose an unreported literature review about clinical findings in patients and their family members with Xq21 deletion syndrome. We finally propose a comprehensive review of HL in contiguous gene syndromes in order to confirm the role of cytogenomic microarray analysis to investigate the etiology of unexplained HL.
Genome-wide CNV analysis uncovers novel pathogenic regions in cohort of five multiplex families with neurodevelopmental disorders.
Structural reorganization of chromosomes by genomic duplications and/or deletions are known as copy number variations (CNVs). Pathogenic and disease susceptible CNVs alter gene dosage and its phenotypic expression that often leads to human genetic diseases including Neurological disorders. CNVs affecting same common genes in multiple neurodevelopmental disorders can better explain the shared clinical and genetic aetiology across brain diseases. Our study presents the novel copy number variations in a cohort of five multiplex consanguineous families with intellectual disability, microcephaly, ASD, epilepsy, and neurological syndromic features. Cytoscan HD microarray suite has revealed genome wide deletions, duplications and LOH regions which are co-segregating in the family members for the rare neurodevelopmental syndromic phenotypes. This study identifies 1q21.1 microduplication, 16p11.2 microduplication, Xp11.22 microduplication, 4p12 microdeletion and Xq21.1 microdeletion that significantly contribute to primary disease onset and its progression for the first time in Pakistani families. Our study has potential impact on the understanding of pathogenic genetic predisposition for appearance of complex and heterogeneous neurodevelopmental disorders with otherwise unexplained syndromic features. Identification of altered gene dosage across the genome is helpful in improved diagnosis, better disease management in day-to-day life activities of patients with cognitive impairment and genetic counselling of families where consanguinity is a tradition. Our study will contribute to expand the knowledge of genotype-phenotype expression and future gateways in therapeutics and precision medicine research will be open in Pakistan.
Intragenic Microdeletion of ULK4 and Partial Microduplication of BRWD3 in Siblings with Neuropsychiatric Features and Obesity.
ULK4 and BRWD3 deletions have been identified in patients with developmental/language delay and intellectual disability. Both genes play pivotal roles in brain development. In particular, ULK4 encodes serine/threonine kinases that are critical for the development and function of the nervous system, while BRWD3 plays a crucial role in ubiquitination, as part of the ubiquitin/proteasome system. We report on 2 brothers, aged 7.6 and 20 years, presenting with cognitive impairment, epilepsy, autistic features, hearing loss, and obesity. Array-CGH analysis demonstrated 2 rare CNVs in both siblings: a paternally inherited microdeletion of ∼145 kb at 3p22.1, disrupting the ULK4 gene, and a maternally inherited microduplication of ∼117 kb at Xq21.1 including only the BRWD3 gene. As already described for other recurrent syndromes with variable phenotype, these findings are challenging in genetic counseling because of an evident variable penetrance. We discuss the possible correlations between the clinical phenotype of our patients and the function of the genes involved in these microrearrangements.
Short stature and primary ovarian insufficiency possibly due to chromosomal position effect in a balanced X;1 translocation.
Primary ovarian insufficiency (POI) is defined as a primary ovarian defect characterized by absent menarche (primary amenorrhea), a decrease in the initial primordial follicle number, high follicle-stimulating hormone (FSH) levels and hypoestrogenism. Although the etiology of a majority of POI cases is not yet identified, several data suggest that POI has a strong genetic component. Conventional cytogenetic and molecular analyses have identified regions of the X chromosome that are associated with ovarian function, as well as POI candidate genes, such as FMR1 and DIAPH2. Here we describe a 10.5-year-old girl presenting with high FSH and luteinizing hormone (LH) levels, pathologic GH stimulation arginine and clonidine tests, short stature, pterygium, ovarian dysgenesis, hirsutism and POI. Cytogenetic analysis demonstrated a balanced reciprocal translocation between the q arms of chromosomes X and 1, with breakpoints falling in Xq21 and 1q41 bands. Molecular studies did not unravel any chromosome microdeletion/microduplication, and no XIST-mediated inactivation was found on the derivative chromosome 1. Interestingly, through immunofluorescence assays, we found that part of the Xq21q22 trait, translocated to chromosome 1q41, was late replicating and therefore possibly inactivated in 30 % metaphases both in lymphocytes and skin fibroblasts, in addition to a skewed 100 % inactivation of the normal X chromosome. These findings suggest that a dysregulation of gene expression might occur in this region. Two genes mapping to the Xq translocated region, namely DIAPH2 and FMR1, were found overexpressed if compared with controls. We report a case in which gonadal dysgenesis and POI are associated with over-expression of DIAPH2 gene and of FMR1 gene in wild type form. We hypothesize that this over-expression is possibly due to a phenomenon known as "chromosomal position effect", which accounts for gene expression variations depending on their localization within the nucleus. For the same effect a double mosaic inactivation of genes mapping to the Xq21-q22 region, demonstrated by immunofluorescence assays, may be the cause of a functional Xq partial monosomy leading to most Turner traits of the proband's phenotype.
Publicações recentes
Contiguous Gene Syndromes and Hearing Loss: A Clinical Report of Xq21 Deletion and Comprehensive Literature Review.
Genome-wide CNV analysis uncovers novel pathogenic regions in cohort of five multiplex families with neurodevelopmental disorders.
Intragenic Microdeletion of ULK4 and Partial Microduplication of BRWD3 in Siblings with Neuropsychiatric Features and Obesity.
Short stature and primary ovarian insufficiency possibly due to chromosomal position effect in a balanced X;1 translocation.
Microdeletion and microduplication 17q21.31 plus an additional CNV, in patients with intellectual disability, identified by array-CGH.
📚 EuropePMCmostrando 4
Contiguous Gene Syndromes and Hearing Loss: A Clinical Report of Xq21 Deletion and Comprehensive Literature Review.
GenesGenome-wide CNV analysis uncovers novel pathogenic regions in cohort of five multiplex families with neurodevelopmental disorders.
HeliyonIntragenic Microdeletion of ULK4 and Partial Microduplication of BRWD3 in Siblings with Neuropsychiatric Features and Obesity.
Cytogenetic and genome researchShort stature and primary ovarian insufficiency possibly due to chromosomal position effect in a balanced X;1 translocation.
Molecular cytogeneticsAssociações
Organizações que acompanham esta doença — pra ter apoio e orientação
Ainda não temos associações cadastradas para Síndrome de microdeleção Xq21.
É de uma associação que acompanha esta doença? Fale com a gente →
Comunidades
Grupos ativos de quem convive com esta doença aqui no Raras
Ainda não existe comunidade no Raras para Síndrome de microdeleção Xq21
Pacientes, familiares e cuidadores se organizam em comunidades pra compartilhar experiências, fazer perguntas e se apoiar. Você pode ser o primeiro.
Tire suas dúvidas
Perguntas, dicas e experiências compartilhadas aqui na página
Participe da discussão
Faça login para postar dúvidas, compartilhar experiências e interagir com especialistas.
Fazer loginDoenças relacionadas
Doenças com sintomas parecidos — ajudam quem ainda está buscando diagnóstico
Referências e fontes
Bases de dados externas citadas neste artigo
Publicações científicas
Artigos indexados no PubMed ligados a esta doença no grafo RarasNet — título, periódico e PMID direto da fonte, sem intermediação de IA.
- Contiguous Gene Syndromes and Hearing Loss: A Clinical Report of Xq21 Deletion and Comprehensive Literature Review.
- Genome-wide CNV analysis uncovers novel pathogenic regions in cohort of five multiplex families with neurodevelopmental disorders.
- Intragenic Microdeletion of ULK4 and Partial Microduplication of BRWD3 in Siblings with Neuropsychiatric Features and Obesity.
- Short stature and primary ovarian insufficiency possibly due to chromosomal position effect in a balanced X;1 translocation.
- Microdeletion and microduplication 17q21.31 plus an additional CNV, in patients with intellectual disability, identified by array-CGH.
Bases de dados e fontes oficiais
Identificadores e referências canônicas usadas para montar este verbete.
- ORPHA:1435(Orphanet)
- OMIM OMIM:303110(OMIM)
- MONDO:0010558(MONDO)
- GARD:369(GARD (NIH))
- Variantes catalogadas(ClinVar)
- Busca completa no PubMed(PubMed)
- Q4831022(Wikidata)
Dados compilados pelo RarasNet a partir de fontes abertas (Orphanet, OMIM, MONDO, PubMed/EuropePMC, ClinicalTrials.gov, DATASUS, PCDT/MS). Este conteúdo é informativo e não substitui avaliação médica.
Conteúdo mantido por Agente Raras · Médicos e pesquisadores podem colaborar